Bayesian multivariate inverse variance weighted model with a choice of prior distributions fitted using RStan.
Source:R/mvmr_ivw_stan.R
mvmr_ivw_stan.RdBayesian multivariate inverse variance weighted model with a choice of prior distributions fitted using RStan.
Usage
mvmr_ivw_stan(
data,
prior = 1,
n.chains = 3,
n.burn = 1000,
n.iter = 5000,
seed = 12345,
...
)Arguments
- data
A data of class
mvmr_format.- prior
An integer for selecting the prior distributions;
1selects a non-informative set of priors;2selects weakly informative priors;3selects a pseudo-horseshoe prior on the causal effect.
- n.chains
Numeric indicating the number of chains used in the HMC estimation in rstan, the default is
3chains.- n.burn
Numeric indicating the burn-in period of the Bayesian HMC estimation. The default is
1000samples.- n.iter
Numeric indicating the number of iterations in the Bayesian MCMC estimation. The default is
5000iterations.- seed
Numeric indicating the random number seed. The default is
12345.- ...
Additional arguments passed through to
rstan::sampling().
Value
An object of class rstan::stanfit.
References
Burgess, S., Butterworth, A., Thompson S.G. Mendelian randomization analysis with multiple genetic variants using summarized data. Genetic Epidemiology, 2013, 37, 7, 658-665 doi:10.1002/gepi.21758 .
Stan Development Team (2020). "RStan: the R interface to Stan." R package version 2.19.3, https://mc-stan.org/.
Examples
if (requireNamespace("rstan", quietly = TRUE)) {
dat <- mvmr_format(
rsid = dodata$rsid,
xbeta = cbind(dodata$ldlcbeta,dodata$hdlcbeta,dodata$tgbeta),
ybeta = dodata$chdbeta,
xse = cbind(dodata$ldlcse,dodata$hdlcse,dodata$tgse),
yse = dodata$chdse
)
suppressWarnings(mvivw_fit <- mvmr_ivw_stan(dat, refresh = 0L))
print(mvivw_fit)
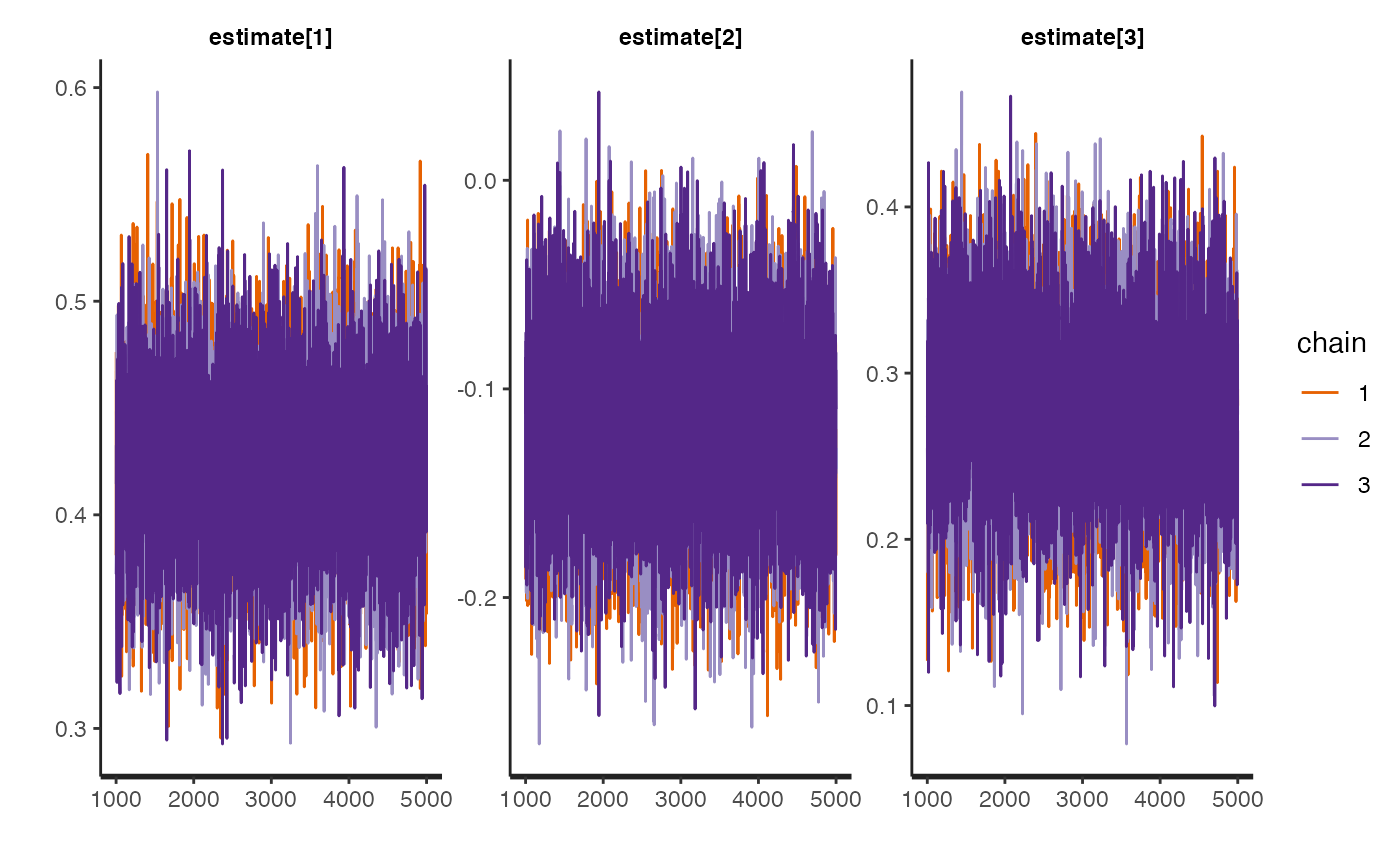
rstan::traceplot(mvivw_fit)
}
#> Inference for Stan model: mvmrivw.
#> 3 chains, each with iter=5000; warmup=1000; thin=1;
#> post-warmup draws per chain=4000, total post-warmup draws=12000.
#>
#> mean se_mean sd 2.5% 25% 50% 75% 97.5% n_eff
#> estimate[1] 0.42 0.00 0.04 0.35 0.40 0.42 0.45 0.50 6505
#> estimate[2] -0.12 0.00 0.04 -0.20 -0.15 -0.12 -0.09 -0.04 6596
#> estimate[3] 0.28 0.00 0.05 0.18 0.25 0.28 0.31 0.38 6174
#> lp__ -198.11 0.02 1.23 -201.39 -198.66 -197.78 -197.21 -196.73 4664
#> Rhat
#> estimate[1] 1
#> estimate[2] 1
#> estimate[3] 1
#> lp__ 1
#>
#> Samples were drawn using NUTS(diag_e) at Mon Aug 19 14:09:26 2024.
#> For each parameter, n_eff is a crude measure of effective sample size,
#> and Rhat is the potential scale reduction factor on split chains (at
#> convergence, Rhat=1).